I am heterozygous for the H63D mutation. Will I get a neurodegenerative disease?
May 9, 2017

- Related Topics:
- Genes to proteins,
- Consumer genetic testing,
- Medical genetics
An undergraduate from Illinois asks:
"If different from Title I found out that I am heterozygous for the H63D hemochromatosis mutation. I’m really worried, because I’ve read that it’s associated with neurodegenerative diseases. Is this true? Does this mean I will get a neurodegenerative disease? Also, is this kind of mutation uncommon?"
Thank you for your question! I’ll try to answer it up front for you, but you can find a full explanation below.
People like you with one copy of H63D (“heterozygotes”) are at a higher risk for certain neurodegenerative diseases. But not by much.
For example, one study on late onset amyotrophic lateral sclerosis (ALS) showed that people like you are around 1.5 times more likely to end up with ALS.1 But this isn’t actually that high of a risk.
Only about 1 in 25,000 Americans has ALS. So your risk would rise to 1 in 16,700 or so.2 Yes, a higher risk, but still extremely unlikely.
And H63D has also been implicated in other neurodegenerative diseases like late onset Alzheimer’s disease (AD).3 But, not every person with H63D ends up with Alzheimer’s
As you can probably guess, having H63D does not mean you will for sure end up with ALS, or another neurodegenerative disease. It just makes it a bit more likely.
This all makes sense when you think about how common H63D is.
According to one study I found, 1 in 5 Caucasian Australians has one copy of H63D.4 Importantly, they do not all have diseases like ALS.

Another study found that over 30% of Americans have one or two copies of either C282Y (a similar mutation) or H63D.5 Again, 30% of Americans do not have a neurodegenerative disease.
So having H63D does not mean you will for sure end up with a neurodegenerative disorder. Not by a long shot.
Which makes sense given the “problems” that H63D can cause.
Iron Problems
H63D is most famous for being involved in something called hereditary hemochromatosis. Basically people with this disease have too much iron in their blood.
Typically, this disease is easily controlled by frequent blood donations that help keep the amount of iron in a persons’ body at a safe level.
But more likely than not, you don’t have to worry about this. See people who are heterozygous for H63D almost never end up with too much iron in their blood.6
Usually you need to also have another mutation, like C282Y, to be at a higher risk for hereditary hemochromatosis.
And even these people don’t get it for sure. They are at a much higher risk, but not everyone with both H63D and C282Y ends up with the disease.7
It might be a bit surprising that H63D turned up as a risk factor for neurodegenerative diseases.
How can a somewhat wonky iron system make getting a neurodegenerative disease like ALS more likely?
Scientists don’t know the answer to this question just yet. But they have a few theories…
One study I found suggested that having the H63D mutation causes extra stress in certain parts of cells. This study thought this extra stress might lead to nerve cells being more prone to being damaged or lost.8
Other scientists think that the H63D might lead to increased iron levels in the brain. This could explain why the mutation is over represented in people with neurodegenerative disorders.9
As you can see, there’s still a lot left to figure out. And unfortunately we don’t know the exact way that having the H63D increases a persons’ risk for certain neurodegenerative disorders.
But the important thing to remember is that even if it does increase someone’s risk, it only seems to do so by a small amount!
What is H63D anyway?
So, that’s your risk for a neurodegenerative disease. Now I want to go into a little more detail about what H63D actually is. And why it causes the problems it does.
To do this, let’s start from the beginning…
The instructions for making humans are found in the 20,000+ genes scattered across our DNA. Each of our genes contains the instructions for one small part of us.
So there is a gene that tells our bodies whether you can drink milk without getting a stomach ache. And one that tells you whether you’ll have red hair or not.
And there are actually a bunch of genes that make sure we have just the right amount of iron in our bodies. This last set of genes includes an important one called HFE.
The HFE gene helps humans control iron levels by detecting the amount of iron in our bodies. It uses this information to control another gene called HAMP.

Very fascinating, but what does this have to do with H63D? Well, H63D describes a very specific mistake, a mutation, in the HFE gene.
The H63D mutation makes it so that HFE can’t do its job, which can cause problems with how we sense and use iron. This is why it makes sense that it can be involved in iron-related diseases like hemochromatosis. (Again, not on its own.)
So that is what H63D is. Sort of. See, the HFE gene doesn’t really do anything on its own.
Instead it has the instructions for the human hemochromatosis protein. And human hemochromatosis protein is the tiny molecular machine that does all the actual iron level sensing.
Genes Have the Instructions for Proteins
Genes have the instructions for proteins, molecules that carry out the functions our bodies need to grow and develop. So the HFE gene has the instructions for the human hemochromatosis protein.
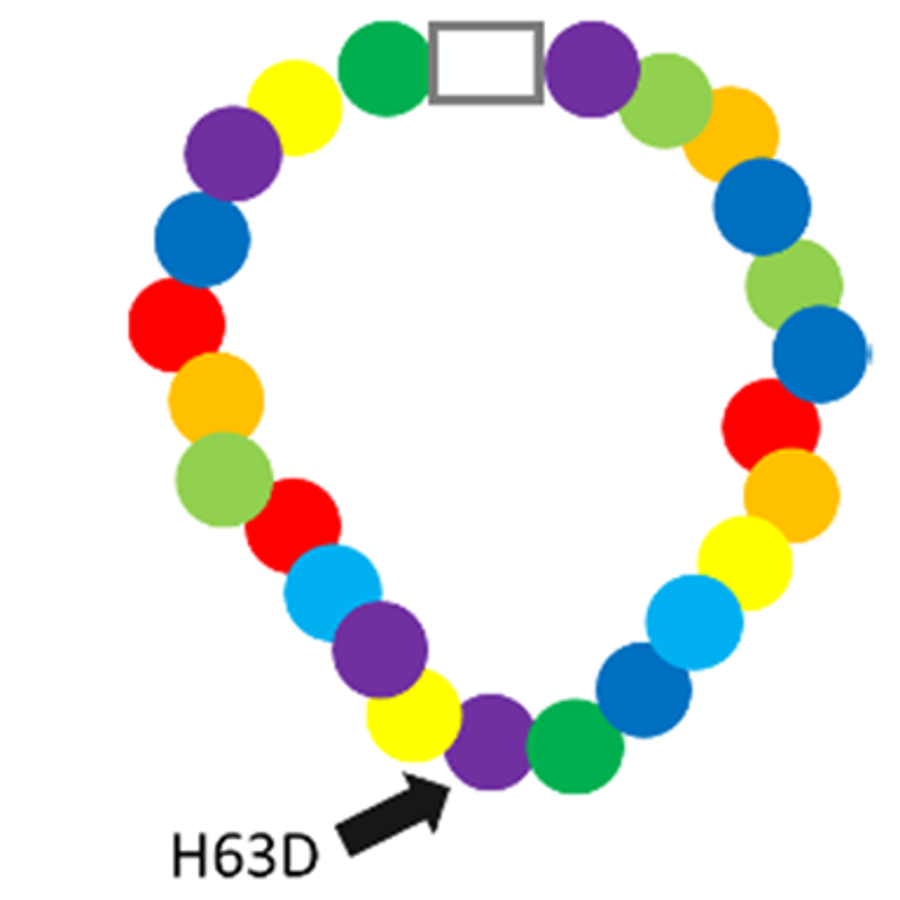
All proteins are made of certain chemicals called amino acids. If we think of amino acids as beads, then a protein is like a necklace. That is, it’s a bunch of beads/amino acids strung together.
Just like there are lots of different styles of beads, there are lots of different types of amino acids - 20 to be exact.
Different beads can be strung together in different orders to make different styles of necklaces. Similarly, different amino acids come together in unique orders to make different proteins.
What the name for the H63D mutation tells us is that normally in this necklace the 63rd bead is the amino acid histidine (H). But, people with this mutation have an aspartic acid (D) bead there instead.
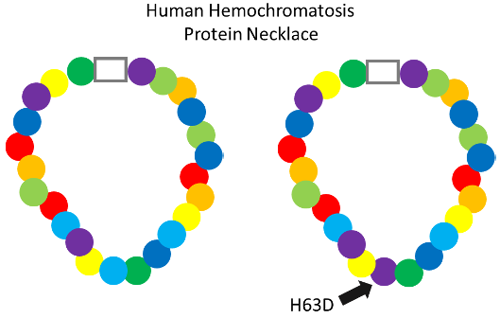
 This change causes this protein to not do its job as well. A single wrong bead has put a kink in the necklace!
This change causes this protein to not do its job as well. A single wrong bead has put a kink in the necklace!
So this is why H63D can cause problems with iron in our bodies. It results in an iron-sensing protein that can’t sense iron as well as the un-mutated version.
Again, we don’t know why this might increase someone’s risk for a neurodegenerative disease. But knowing what the HFE gene does and how the H63D mutation affects it helps point us in the right direction to figure it out.
Read More:
- Medline: Hereditary hemochromatosis
- Haemochromatosis Australia: Genetics of haemochromatosis

Author: Megan Nathan
When this answer was published in 2017, Megan was a student in the Stanford MS Program in Human Genetics and Genetic Counseling. Megan wrote this answer while participating in the Stanford at The Tech program.
 Skip Navigation
Skip Navigation
