
How do things move around inside a cell?
December 19, 2019

- Related Topics:
- Molecular biology,
- Genes to proteins,
- Transcription and translation,
- RNA
A curious adult from Arkansas asks:
"I am trying to find out about movement/propulsion in the cell. More specifically:
- During transcription, what moves or propels RNA polymerase along the DNA template strand to create the mRNA?
- What propels and guides the mRNA from the nucleus through the nuclear pore and into the cytoplasm?
- During translation, what propels the ribosome along the mRNA as it assembles the polypeptide chain?"
There are several strategies our cells use to move (literally) through transcription and translation. Before we get to it, let’s first paint ourselves a picture of what our cells look like.
We learn in intro biology classes about animal cells. Usually, we’re shown a picture like the one below:
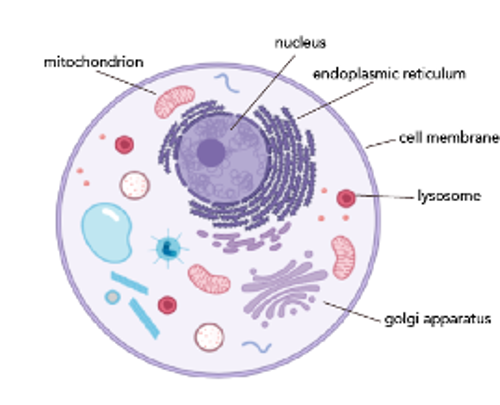
While this diagram is great for helping us get a sense of what the major organelles are, it does a disservice by making us think there’s this large pool of unoccupied space in both the cytoplasm and nucleus. In truth, our cells have no empty space!
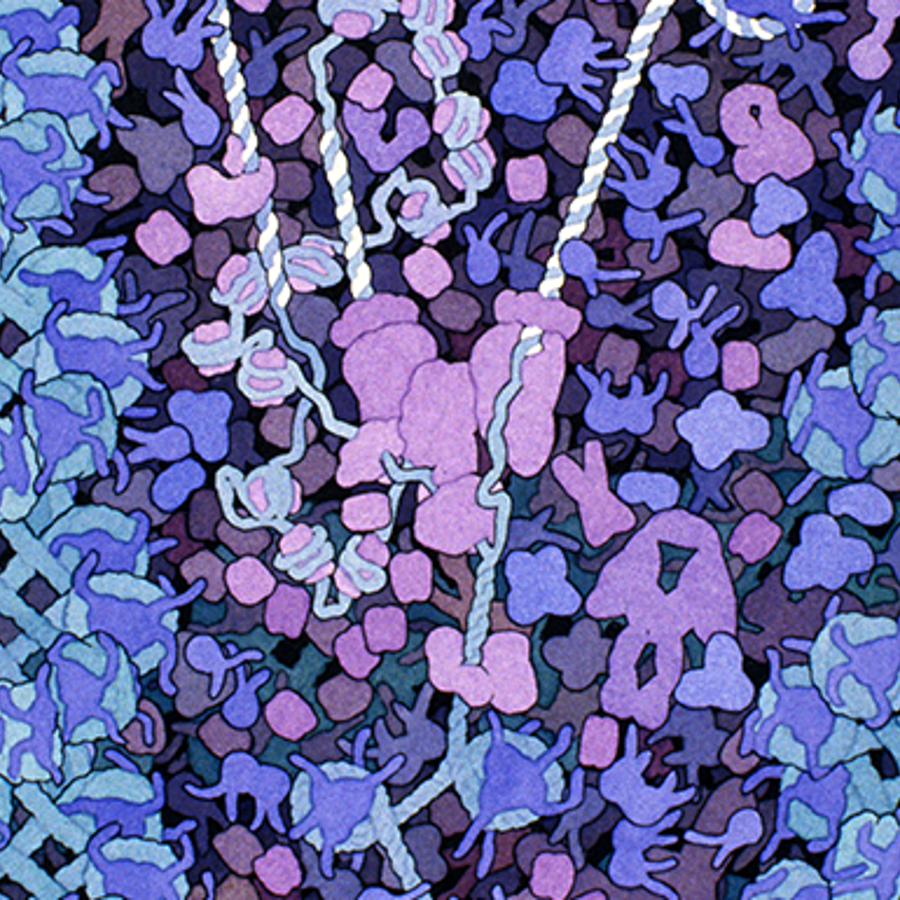
The cell is a very crowded place. Chock full of macromolecules, such as enzymes, filamentous proteins, and organelles all suspended in water. Take a look at this artistic interpretation of the nucleus:
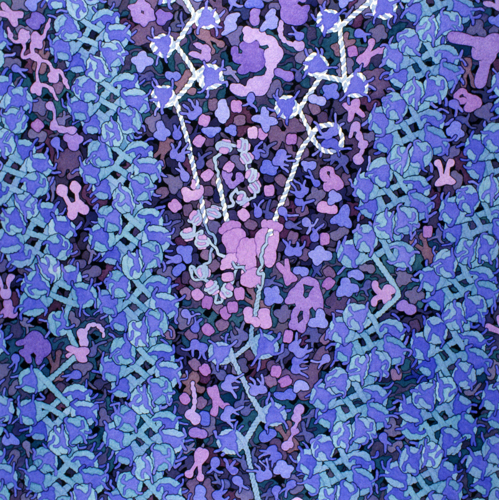
This is actually pretty close to what microscopy images look like!
So how do any of these molecules find their destination in such a crowded space?
Random Motion
Our cells are a packed party of wiggling and jiggling proteins always bumping into one another. Because of this constant motion, it’s only a matter of time before two proteins find one another.
All of these molecules shown above are in constant motion. Fast-moving water molecules collide into the larger macromolecules causing them to “dance” around in random motion.
This random motion was first observed in 1827 by Robert Brown. While examining pollen grains suspended in water, he noticed the pollen jittering around in the petri dish. Today we call this type of movement brownian motion.
This kind of movement is entirely random. There’s no preferred direction. Molecules just bump, dance, and wiggle around until they get to their destination.
So the first answer to all three of your questions is there’s actually constant random movements!
But not all motion in the cell is random! How do those motions happen?
Molecular guides
As you mention, after the mRNA is made in the nucleus, it exits out into the cytoplasm via the nuclear pore.
But how does an mRNA get to the nuclear pore?
The mRNA would make it to the nuclear pore eventually from brownian motion. But what if we need it now? Are we going to wait all day until it randomly finds the exit?
It turns out that there are some helper proteins in the nucleus. They bind to the mRNA and direct it to an exit point.
These helper proteins are usually made up of two important parts. One part grabs on to the RNA. The other part binds to the nuclear pore. Together, these two parts act like a magnet that guides the mRNA exactly where it needs to go.
Here’s what one such helper protein, NFX1, looks like:
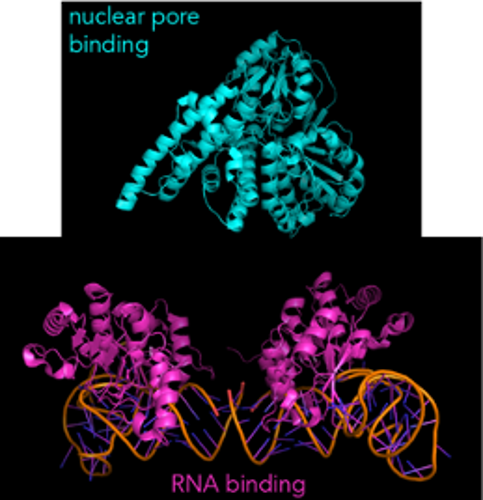
(As a small aside, NFX1 is actually one large protein with each domain connected. But because it’s often difficult to get the structure of large proteins, researchers will make smaller fragments of the protein and get separate structures for each fragment.)
This molecular guide helps mRNA exit the nucleus quickly. Much faster than if it had to rely on random motion!
Proteins as machines
Ok so helper proteins can guide molecules from Point A to Point B. But how do molecules move along a different molecule? What causes polymerase to travel across a strand of DNA, or ribosomes to move forward along an mRNA?
These are actually still really big areas of research. We don’t have all the answers yet, but we do know a bit about how these work.
Like the helper proteins mentioned before, ribosomes and polymerases can grab on to other molecules. But they do so much more than passively hold on! They can also do chemical reactions. This makes them a special type of protein, called enzymes.
Enzymes are able to actually change molecules into different things. The molecule it starts with is called the substrate. The end result is called the product.
There are a few different ways that enzymes can do this, but we’ll just focus on one method here.
Sometimes an enzyme needs to change shape to grab the molecule it’s working on. The substrate might sort of fit initially, but the enzyme needs to readjust to get that perfect fit. Sometimes these movements are small. But sometimes the enzyme completely rearranges! This is called induced fit binding. Here’s what that looks like:
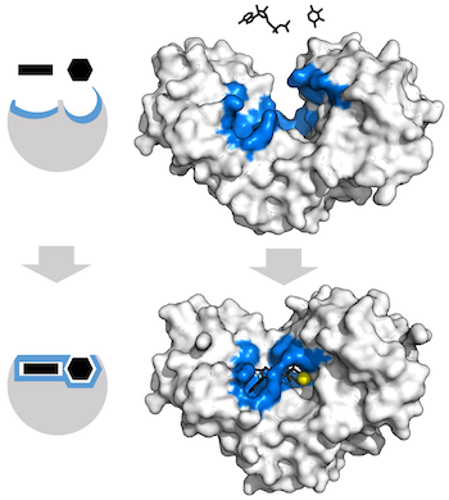
As polymerases and ribosomes do their work, they go through a few different shape changes. These changes help make them move in the correct direction.
For example, the ribosome changes shape quite a bit during translocation. Each time it adds a tRNA to the polypeptide chain, it rearranges.
The image below shows just one part of the ribosome complex. Take a look at how much that light blue domain on the bottom sprawls out!
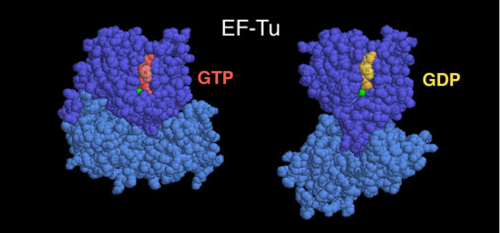
Something else happens when the ribosome changes shape: it inches forward. This motion is no longer random. It is now directional.
You might ask: well, why does it move forward instead of backwards?
This was a big area of research over the last decade, with lots of scientists investigating this very question. What has emerged from that research is that these enzymes behave as molecular ratchets.
You might have used a socket ratchet before, and know it can only turn in one direction. You can only turn it one way because the teeth get stuck in the other direction:
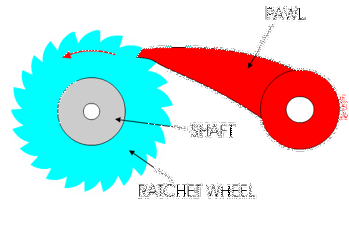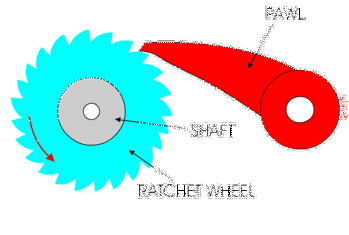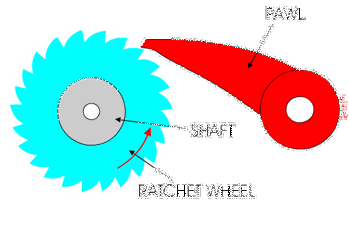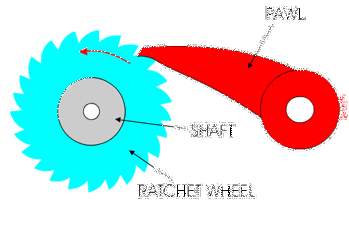
Ribosomes and polymerases alternate between two shapes. They rock back and forth between the two conformations.
Each time the enzyme is in one shape, it has the opportunity to move forward or backwards. But when it moves forward, another shape change occurs that blocks the backward movement.
These ratchet-like movements of the enzyme are what allow it to move forward along the template and quickly proceed through transcription and translation!
Read More:
-
TED talk: Animations of unseeable biology
-
More molecular art by David Goodsell can be found on his website

Author: Nicole DelRosso
When this answer was published in 2019, Nicole was a Ph.D. candidate in the Department of Biophysics, studying high throughput biophysical screens of transcription factors in both Polly Fordyce’s & Lacra Bintu’s laboratories. She wrote this answer while participating in the Stanford at The Tech program.
 Skip Navigation
Skip Navigation
