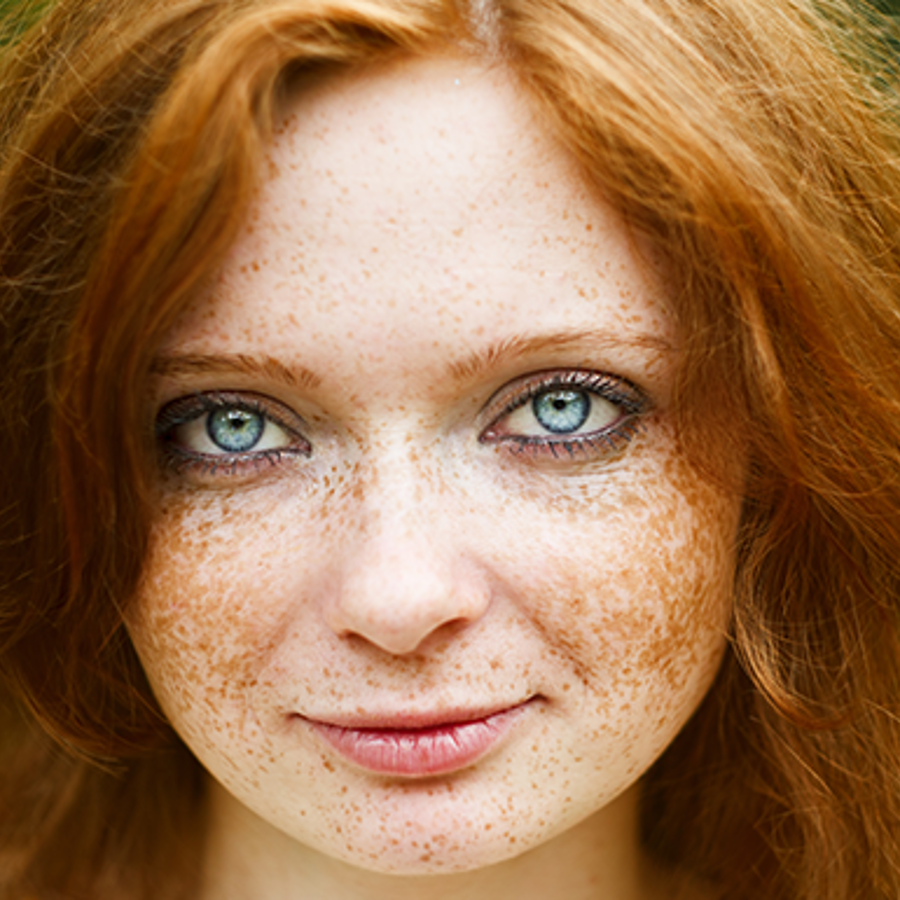

Do recessive genes not make any proteins? What exactly do they do?
March 27, 2013

- Related Topics:
- Molecular biology,
- Dominant and recessive,
- Classical genetics
A curious adult from the US asks:
"In people with one dominant and one recessive gene, does the recessive gene not make anything? If so, does the dominant gene stop the recessive gene from making anything? Or, does the recessive gene make a broken protein? If the recessive gene makes something, shouldn’t it be incompletely dominant or co-dominant?"
Words like dominant and recessive don’t describe the gene…they describe the trait.
What this means is that whether a particular version of a gene is actually doing something doesn’t matter in terms of the trait being recessive or not. What matters is what trait we are looking at.
This can mean that the same version (or allele) of a gene can be dominant, recessive or even co-dominant. It all depends on the trait. A great example of this is broken versions of the MC1R gene.
The MC1R gene is a key player in both red hair and freckles. Red hair is a recessive trait meaning you need two broken MC1R copies to end up with it. But having freckles, which are caused by the same broken MC1R genes, is a dominant trait. One copy is enough. So in this case, the same allele gives a different result depending on the trait we focus on.
Another way of putting this is that your question really boils down to the difference between genotype and phenotype. Genotype is the genes that we have and phenotype is the trait those genes give us.
All of the terms you’ve mentioned—dominant, recessive, co-dominant and incomplete dominance—have to do with what trait appears in a living thing. They have to do with phenotype. None of them has anything to do with the genotype (although as you’ll see, the two are definitely related!).
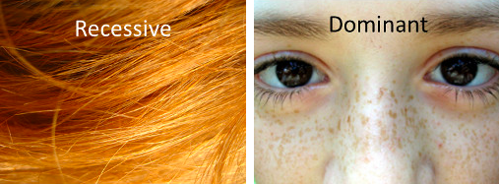
The Many Ways to Make a Recessive Allele
A gene has the instructions for making a specific protein. And each protein does a certain job in the cell.
So let’s say there is a gene involved in making a flower red. It makes a protein whose job it is to make a red pigment.
This gene comes in two forms (or alleles), R and r. This flower, like us, has two copies of each of its genes. So in terms of our red flower gene, our flower can be RR, Rr or rr.
Let’s say that both RR and Rr are red and rr is white. If this is the case, then R is dominant over r because RR and Rr are the same color. We can also say that r is recessive to R for the same reason.
Now it won’t matter if r is recessive because it makes no protein, a broken protein or a version of the protein that makes white pigment. In each case, r is recessive to R.
Let’s say instead that RR is red, Rr is pink and rr is white. When this happens, we say that R is incompletely dominant over r. But again, the reason why being Rr leads to a pink color doesn’t matter in terms of this being incompletely dominant.
One way Rr might end up pink is something called haploinsufficiency. The idea here is that one R only makes enough pigment to give a pink color and so you need both to be R to get red. In this case, r might make no protein or a defective one and you’d still get pink.
Another way incomplete dominance might work is if R makes a protein that makes red pigment and r makes one that makes a white pigment. Both make a working protein but the white dilutes the red leading to pink.
Rr could also be a flower that is both red and white. This is called co-dominance. Co-dominance is when you see both traits at once.
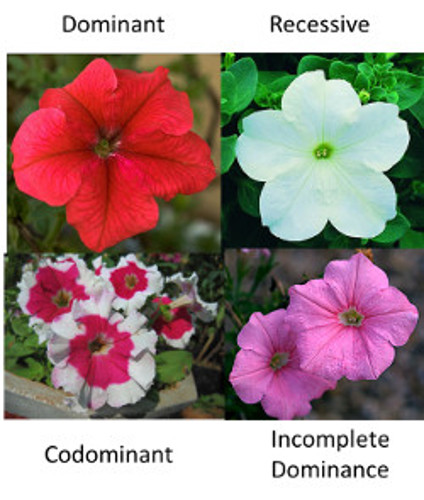

As you can see, the same effects on a gene can lead to different traits. The r allele that makes no protein could lead to a recessive phenotype if the red allele makes a lot of red pigment. Or the same r allele could lead to incomplete dominance if the red allele only makes enough pigment to make the flower pink.
The same thing is true with an r allele that makes a white pigment. It could be that the red allele makes enough red pigment to overpower the white in which case white is a recessive trait. Or the white could dilute the red and we’d have incomplete dominance.
The reason the combination of genes leads to a certain trait does not affect what we call the trait. So the complete absence of a product from a gene can lead to a recessive allele or an incompletely dominant one. And a functional protein can sometimes be recessive!
The Same Allele can be Dominant, Recessive or Co-dominant.
As I showed with the red flower example, the amount or type of protein the recessive gene makes only matters if it affects the trait. Recessive, dominant, co-dominant, and incomplete dominance all refer to the phenotype, not the genotype.
There is a great real life human example of this in the sickle cell version of the hemoglobin gene. As you can see below, depending on the trait we look at, this same allele can be dominant, recessive or co-dominant.
|
Phenotype |
Dominance |
Genotype |
|
Anemia |
Recessive |
Two copies to become anemic |
|
Malaria resistant |
Dominant |
One copy to become resistant |
|
Blood cell shape |
Co-dominant |
One copy to have a mixture of normal and sickle-cell shaped cells |
Sickle cell anemia is a disease where blood cells sometimes become elongated and curved, like the sickle in the picture below:
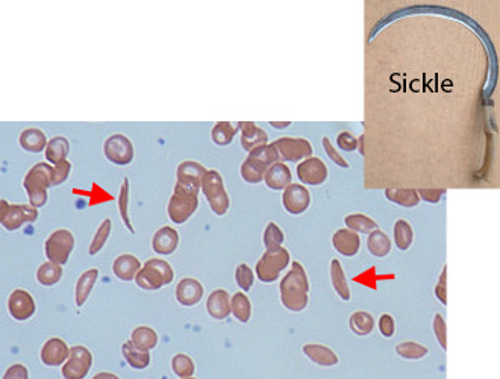
Sickle shaped cells have trouble carrying oxygen, making the person anemic. This happens if both copies of your hemoglobin gene are the sickle shape allele causing all your blood cells to be sickle shaped too.
If just one copy is the sickle cell version, then you are resistant to malaria. You have a mix of sickle shaped and normal doughnut shaped cells. (Click here to learn why this makes someone resistant to malaria.)
So if we look at the shape of blood cells, the sickle cell allele is codominant. And if we look at malaria resistance, it is dominant. And finally, if we look at sickle cell anemia, it is recessive. One allele, three different dominances.
In fact, we can even add a fourth. Malaria resistance can be called overdominant in places where there is a lot of malaria. Overdominance is when the heterozygous phenotype is more advantageous than either of the homozygous phenotypes.
Conclusion
As you can see, when talking about dominant and recessive, we need to focus on the trait and not the gene. They describe phenotype not genotype.
There are many different genotypes that can make a phenotype. Also, the same genotype can make many different phenotypes. So, if you want to understand how a trait (phenotype) is made (genotype), you have to think about what the alleles do in that specific case.
Appendix: Alleles Sometimes Affect Each Other
You also asked whether the protein from one allele could affect protein production from the other allele. In theory, this could happen, but is very uncommon. So uncommon that I cannot think of an example!
However, there are ways for a defective protein to actually be dominant. One way is something called a dominant negative.
In this case, the protein from one allele does not work properly, and causes protein from the other allele to also not work. This can happen when two of the same proteins physically interact in a cell. Red hair is a trait that is thought to sometimes be caused by a dominant negative.
As you might remember from the introduction, MC1R is the key gene involved in red hair. This gene makes a protein whose job it is to turn red pigment into brown. When this protein isn’t doing its job, you end up with a buildup of red pigment and so red hair.
Red hair is usually a recessive trait. You need two broken versions of the MC1R gene to cause the buildup of red pigment. But on rare occasions, red hair can actually be dominant.
Two MC1R proteins need to stick together to change red pigment into brown. Sometimes when a defective version sticks to a working one, the protein can’t do its job. Sort of like a pair of scissors where one blade is defective.
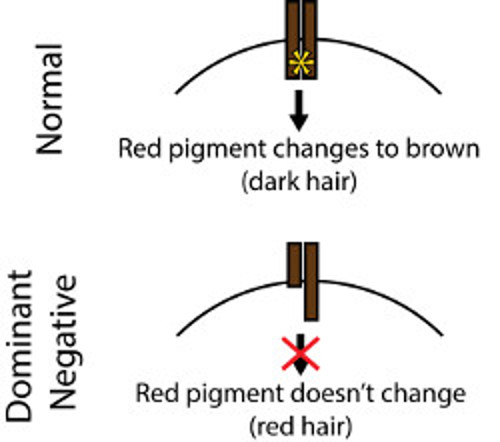
In this case, the defective MC1R protein is actually dominant. You need just one version of it to have red hair. So, what you think would be recessive (a defective protein) is actually dominant!
This is a slightly different twist on what we’ve been talking about. Before we mostly focused on different traits and how their phenotype is affected by the same allele. Here we give an example of how a single trait can be affected by different gene versions. The same trait, red hair, is either dominant or recessive depending on the gene version that caused it!

Author: Dr. Roberta Hannibal
When this answer was published in 2013, Roberta was a post-doctoral fellow in the Department of Genetics, studying placenta evolution, development and genomics in Julie Baker’s laboratory. She wrote this answer while participating in the Stanford at The Tech program.
 Skip Navigation
Skip Navigation
